Reactome Analysis (Robertson Lab)
Ha Tran
22/08/2021
Last updated: 2025-11-27
Checks: 7 0
Knit directory: 5_gd_Tcell/1_analysis/
This reproducible R Markdown analysis was created with workflowr (version 1.7.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(12345) was run prior to running the
code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version b9f184a. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Untracked files:
Untracked: .DS_Store
Untracked: figure/scatter2-1.png
Untracked: figure/scatter3-1.png
Untracked: figure/scatter4-1.png
Untracked: figure/scatter5-1.png
Untracked: figure/scatter_3d4-1.png
Untracked: figure/scatter_interactive4-1.png
Untracked: figure/treemap2-1.png
Untracked: figure/treemap3-1.png
Untracked: figure/treemap4-1.png
Untracked: figure/treemap5-1.png
Unstaged changes:
Modified: 0_data/RDS_plots/go_combined_dotPlot.rds
Modified: 0_data/RDS_plots/go_combined_parTerm_dotPlot.rds
Modified: 0_data/RDS_plots/go_dotPlot.rds
Modified: 0_data/RDS_plots/go_parTerm_dotPlot.rds
Modified: 0_data/RDS_plots/go_parTerm_scatter.rds
Modified: 0_data/RDS_plots/kegg_dotPlot.rds
Modified: 0_data/RDS_plots/kegg_path_Hmap.rds
Modified: 0_data/RDS_plots/ma_plots.rds
Modified: 0_data/RDS_plots/react_combined_dotPlot.rds
Modified: 0_data/RDS_plots/react_dotPlot.rds
Modified: 0_data/RDS_plots/vol_plots.rds
Modified: 2_plots/1_QC/PC1_PC2.svg
Modified: 2_plots/1_QC/PC1_PC3.svg
Modified: 2_plots/1_QC/PC2_PC3.svg
Modified: 2_plots/2_DE/heat_down_INT vs CONT.svg
Modified: 2_plots/2_DE/heat_down_INT vs SVX_VAS.svg
Modified: 2_plots/2_DE/heat_down_INT vs VAS.svg
Modified: 2_plots/2_DE/heat_down_SVX vs SVX_VAS.svg
Modified: 2_plots/2_DE/heat_down_SVX_VAS vs CONT.svg
Modified: 2_plots/2_DE/heat_down_VAS vs SVX_VAS.svg
Modified: 2_plots/2_DE/heat_up_INT vs CONT.svg
Modified: 2_plots/2_DE/heat_up_INT vs SVX_VAS.svg
Modified: 2_plots/2_DE/heat_up_INT vs VAS.svg
Modified: 2_plots/2_DE/heat_up_SVX vs SVX_VAS.svg
Modified: 2_plots/2_DE/heat_up_SVX_VAS vs CONT.svg
Modified: 2_plots/2_DE/heat_up_VAS vs SVX_VAS.svg
Modified: 2_plots/2_DE/hist_INT vs CONT.svg
Modified: 2_plots/2_DE/hist_INT vs SVX_VAS.svg
Modified: 2_plots/2_DE/hist_INT vs VAS.svg
Modified: 2_plots/2_DE/hist_SVX vs SVX_VAS.svg
Modified: 2_plots/2_DE/hist_SVX_VAS vs CONT.svg
Modified: 2_plots/2_DE/hist_VAS vs SVX_VAS.svg
Modified: 2_plots/3_FA/go/combine_go_dot.svg
Modified: 2_plots/3_FA/go/parTerm_dot_INT vs CONT.svg
Modified: 2_plots/3_FA/go/parTerm_dot_INT vs SVX_VAS.svg
Modified: 2_plots/3_FA/go/parTerm_dot_SVX vs SVX_VAS.svg
Modified: 2_plots/3_FA/go/parTerm_dot_SVX_VAS vs CONT.svg
Modified: 2_plots/3_FA/go/parTerm_dot_VAS vs SVX_VAS.svg
Modified: 2_plots/3_FA/go/semSim_dendrogram_INT vs CONT.svg
Modified: 2_plots/3_FA/go/semSim_dendrogram_INT vs SVX_VAS.svg
Modified: 2_plots/3_FA/go/semSim_dendrogram_SVX vs SVX_VAS.svg
Modified: 2_plots/3_FA/go/semSim_dendrogram_SVX_VAS vs CONT.svg
Modified: 2_plots/3_FA/go/semSim_dendrogram_VAS vs SVX_VAS.svg
Modified: 2_plots/3_FA/go/semSim_scatter_INT vs CONT.svg
Modified: 2_plots/3_FA/go/semSim_scatter_INT vs SVX_VAS.svg
Modified: 2_plots/3_FA/go/semSim_scatter_SVX vs SVX_VAS.svg
Modified: 2_plots/3_FA/go/semSim_scatter_SVX_VAS vs CONT.svg
Modified: 2_plots/3_FA/kegg/combine_kegg_dot.svg
Modified: 2_plots/3_FA/kegg/heat_Neutrophil extracellular trap formation.svg
Modified: 2_plots/3_FA/kegg/heat_PD-L1 expression and PD-1 checkpoint pathway in cancer.svg
Modified: 2_plots/3_FA/kegg/heat_T cell receptor signaling pathway.svg
Modified: 2_plots/3_FA/kegg/heat_Th1 and Th2 cell differentiation.svg
Modified: 2_plots/3_FA/kegg/heat_Th17 cell differentiation.svg
Modified: 2_plots/3_FA/kegg/kegg_dot_INT vs CONT.svg
Modified: 2_plots/3_FA/kegg/kegg_dot_INT vs SVX_VAS.svg
Modified: 2_plots/3_FA/kegg/kegg_dot_SVX_VAS vs CONT.svg
Modified: 2_plots/3_FA/kegg/kegg_dot_VAS vs SVX_VAS.svg
Modified: 2_plots/3_FA/kegg/kegg_upset_INT vs CONT.svg
Modified: 2_plots/3_FA/kegg/kegg_upset_INT vs SVX_VAS.svg
Modified: 2_plots/3_FA/kegg/kegg_upset_SVX_VAS vs CONT.svg
Modified: 2_plots/3_FA/kegg/kegg_upset_VAS vs SVX_VAS.svg
Modified: 2_plots/3_FA/reactome/combine_react_dot.svg
Modified: 2_plots/3_FA/reactome/react_dot_INT vs CONT.svg
Modified: 2_plots/3_FA/reactome/react_dot_INT vs SVX_VAS.svg
Modified: 2_plots/3_FA/reactome/react_dot_INT vs VAS.svg
Modified: 2_plots/3_FA/reactome/react_dot_SVX vs SVX_VAS.svg
Modified: 2_plots/3_FA/reactome/react_dot_SVX_VAS vs CONT.svg
Modified: 2_plots/3_FA/reactome/react_dot_VAS vs SVX_VAS.svg
Modified: 2_plots/3_FA/reactome/react_upset_INT vs CONT.svg
Modified: 2_plots/3_FA/reactome/react_upset_INT vs SVX_VAS.svg
Modified: 2_plots/3_FA/reactome/react_upset_INT vs VAS.svg
Modified: 2_plots/3_FA/reactome/react_upset_SVX vs SVX_VAS.svg
Modified: 2_plots/3_FA/reactome/react_upset_SVX_VAS vs CONT.svg
Modified: 2_plots/3_FA/reactome/react_upset_VAS vs SVX_VAS.svg
Modified: 2_plots/4_paper/subset_degs.svg
Modified: 2_plots/combine_ipa_dot.svg
Modified: 2_plots/dnf_plot.svg
Modified: 2_plots/intVsvxVAS.svg
Modified: 2_plots/upstream_hmap.svg
Modified: 3_output/DEGs.xlsx
Modified: 3_output/Gene Ontology.xlsx
Modified: 3_output/KEGG.xlsx
Modified: 3_output/Reactome.xlsx
Modified: 3_output/deg_all_new.xlsx
Modified: 3_output/deg_sig_new.xlsx
Modified: 3_output/eigenvalues.xlsx
Modified: 3_output/enrichKEGG_sig.xlsx
Modified: 3_output/reactome_all_new.xlsx
Modified: 3_output/reactome_sig_new.xlsx
Modified: 3_output/semSim_GO_sig.xlsx
Modified: README.html
Modified: README.md
Modified: figure/dot2-1.png
Modified: figure/dot3-1.png
Modified: figure/dot4-1.png
Modified: figure/dot5-1.png
Modified: figure/upset2-1.png
Modified: figure/upset3-1.png
Modified: figure/upset4-1.png
Modified: figure/upset5-1.png
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown (1_analysis/reactome.Rmd) and HTML
(docs/reactome.html) files. If you’ve configured a remote
Git repository (see ?wflow_git_remote), click on the
hyperlinks in the table below to view the files as they were in that
past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| html | d519e7f | Ha Tran | 2024-12-03 | Build site. |
| Rmd | 9fc0156 | Ha Tran | 2024-12-03 | workflowr::wflow_publish(here::here("1_analysis/*.Rmd")) |
| html | 5dce909 | Ha Tran | 2024-11-07 | Build site. |
| html | d71eeb4 | Ha Tran | 2024-10-16 | Build site. |
| Rmd | 5f0c7a1 | Ha Tran | 2024-10-16 | workflowr::wflow_publish(here::here("1_analysis/*.Rmd")) |
| html | ae93fcc | tranmanhha135 | 2024-02-02 | Build site. |
| Rmd | fbcdd69 | tranmanhha135 | 2024-02-02 | workflowr::wflow_publish(here::here("1_analysis/*.Rmd")) |
| html | 2c24612 | tranmanhha135 | 2022-10-13 | build website |
| Rmd | 324032b | tranmanhha135 | 2022-10-11 | resize images |
| html | 11a5cf4 | tranmanhha135 | 2022-10-03 | build wedsite |
| Rmd | 1101367 | tranmanhha135 | 2022-10-02 | Completed functional enrichment for all comparison |
| html | 1101367 | tranmanhha135 | 2022-10-02 | Completed functional enrichment for all comparison |
| html | 68585df | tranmanhha135 | 2022-09-20 | build webste |
| Rmd | 192d010 | tranmanhha135 | 2022-09-20 | functional enrichment with new dataset |
| html | 192d010 | tranmanhha135 | 2022-09-20 | functional enrichment with new dataset |
| Rmd | 0df047f | tranmanhha135 | 2022-09-08 | minor changes to build and publish |
| html | 0df047f | tranmanhha135 | 2022-09-08 | minor changes to build and publish |
| html | 54e0166 | Ha Manh Tran | 2022-01-01 | Build site. |
| html | 32454d5 | Ha Manh Tran | 2022-01-01 | Build site. |
| Rmd | c667dd0 | Ha Manh Tran | 2022-01-01 | workflowr::wflow_publish(files = here::here(c("1_analysis/index.Rmd", |
| Rmd | 3b268fc | Ha Tran | 2021-12-09 | multiple FC, reactome, big clean up. |
| html | 3b268fc | Ha Tran | 2021-12-09 | multiple FC, reactome, big clean up. |
| Rmd | 7247e9c | Ha Tran | 2021-11-30 | additional KEGG heatmap, pathview, reactome |
| html | 7247e9c | Ha Tran | 2021-11-30 | additional KEGG heatmap, pathview, reactome |
# working with data
library(dplyr)
library(magrittr)
library(readr)
library(tibble)
library(reshape2)
library(tidyverse)
library(VennDiagram)
# Visualisation:
library(kableExtra)
library(ggplot2)
library(grid)
library(pander)
library(cowplot)
library(pheatmap)
library(DT)
library(extrafont)
# Custom ggplot
library(ggbiplot)
library(ggrepel)
# Bioconductor packages:
library(edgeR)
library(limma)
library(Glimma)
library(clusterProfiler)
library(org.Mm.eg.db)
library(enrichplot)
library(ReactomePA)
library(pandoc)
library(knitr)
opts_knit$set(progress = FALSE, verbose = FALSE)
opts_chunk$set(warning=FALSE, message=FALSE, echo=FALSE)Reactome
Reactome database provides curated information about biological pathways, including molecular events and reactions within cells. It focuses on human biology and is widely used for pathway analysis and functional interpretation of high-throughput data.
KEGG and Reactome both include approximately the same number of genes. The difference lies in KEGG’s use of broader terms, while Reactome employs similar terms but with multiple detailed entries.
In the Reactome database, terms are organized hierarchically based on the classification of biological pathways. The organization follows a tree-like structure, where terms represent different levels of granularity in understanding molecular events and reactions within cells
Visualisation
The following visualisations are Reactome enrichment analysis performed with set of DE genes. IMPORTANTLY, these Reactome terms are significantly enriched with P value < 0.05.
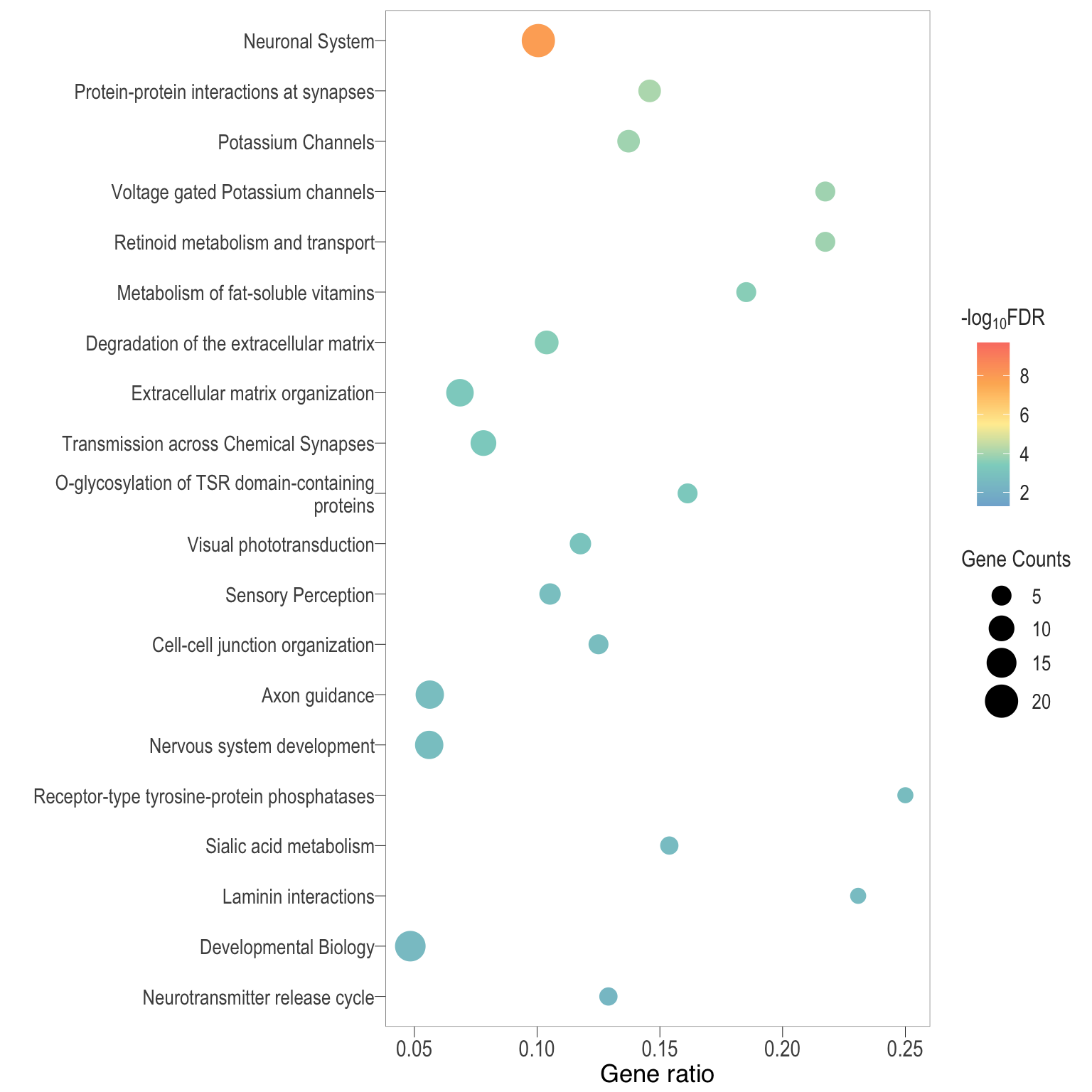
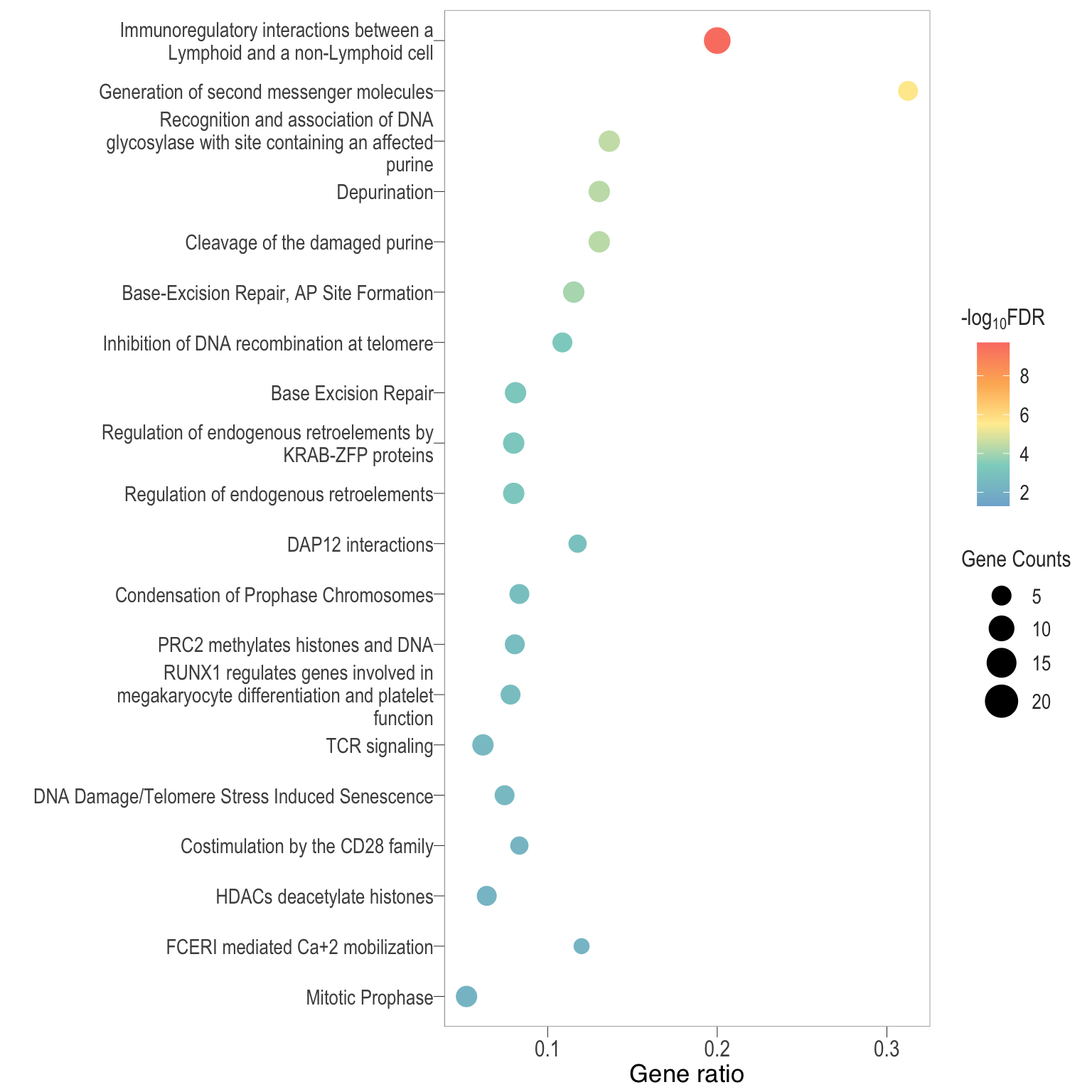
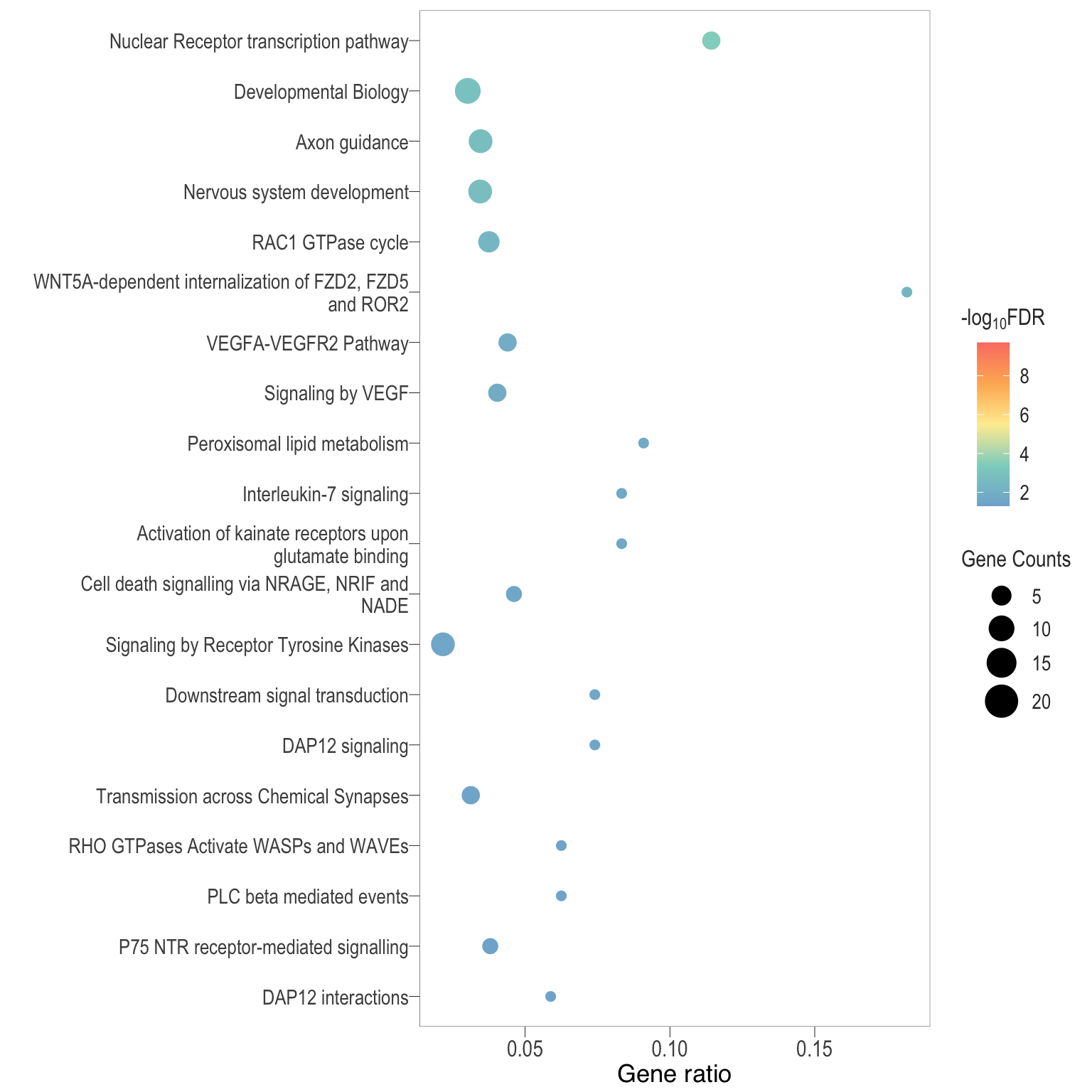
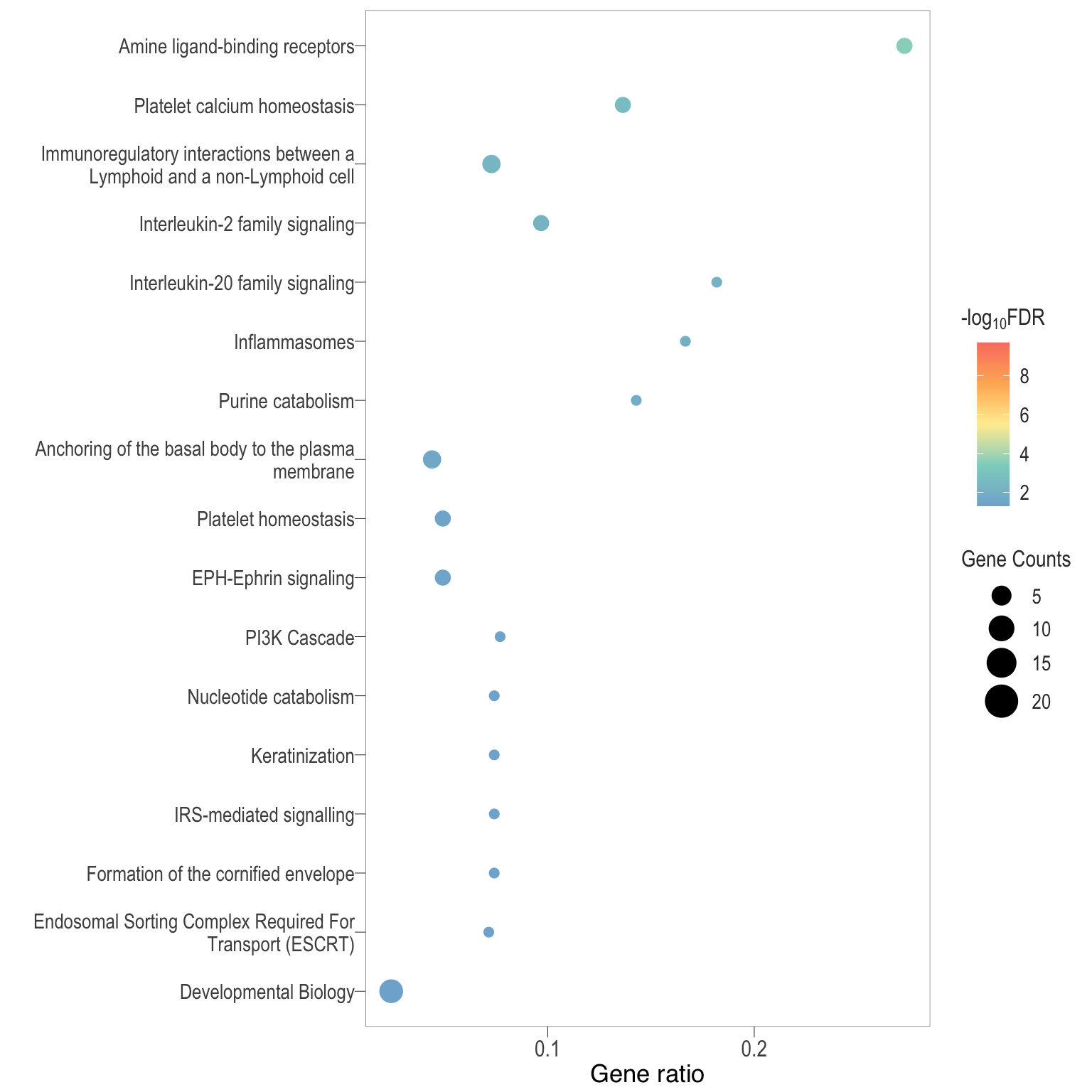
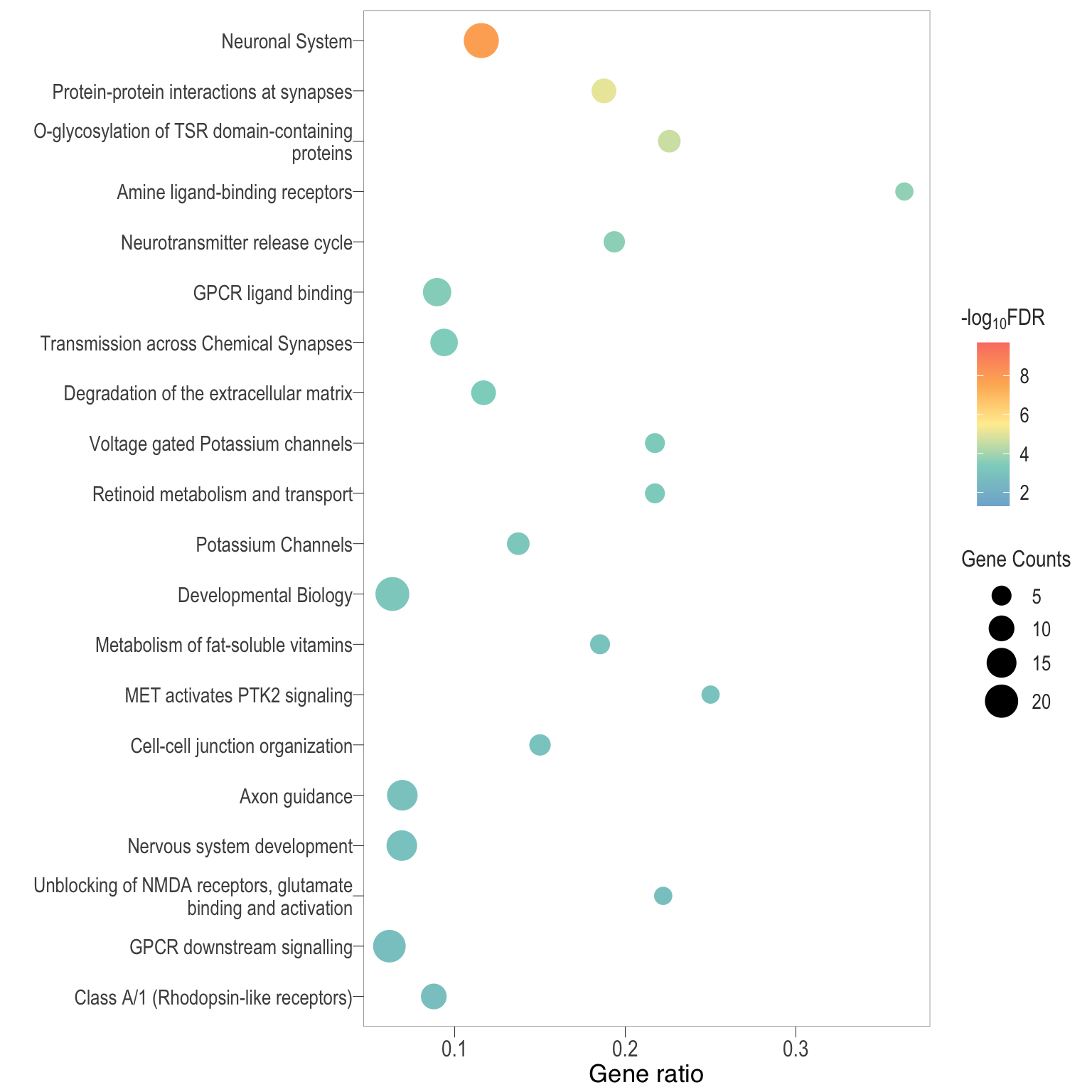
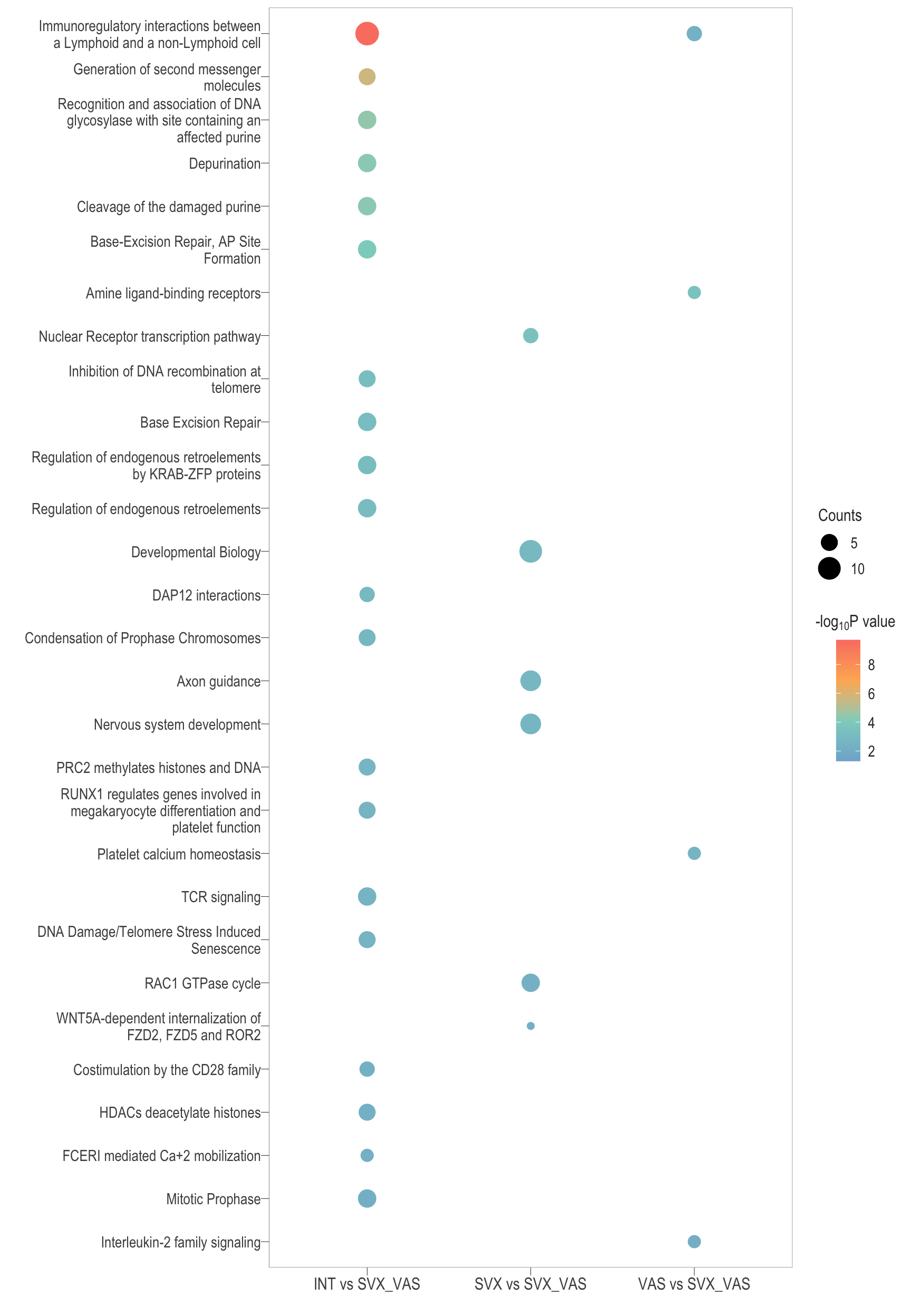
Dot plot: illustrates the top enriched Reactome pathways
- \(Gene ratio =\) the number of significant DE gene in the term / the total of number of genes in the term as indicated by the size
Table: list of all the significant Reactome pathways
- NOTE: To keep this a readable table, the full pathway description were removed, check the exported Excel spreadsheet for full details on pathways class, descriptions, related pathways, and references
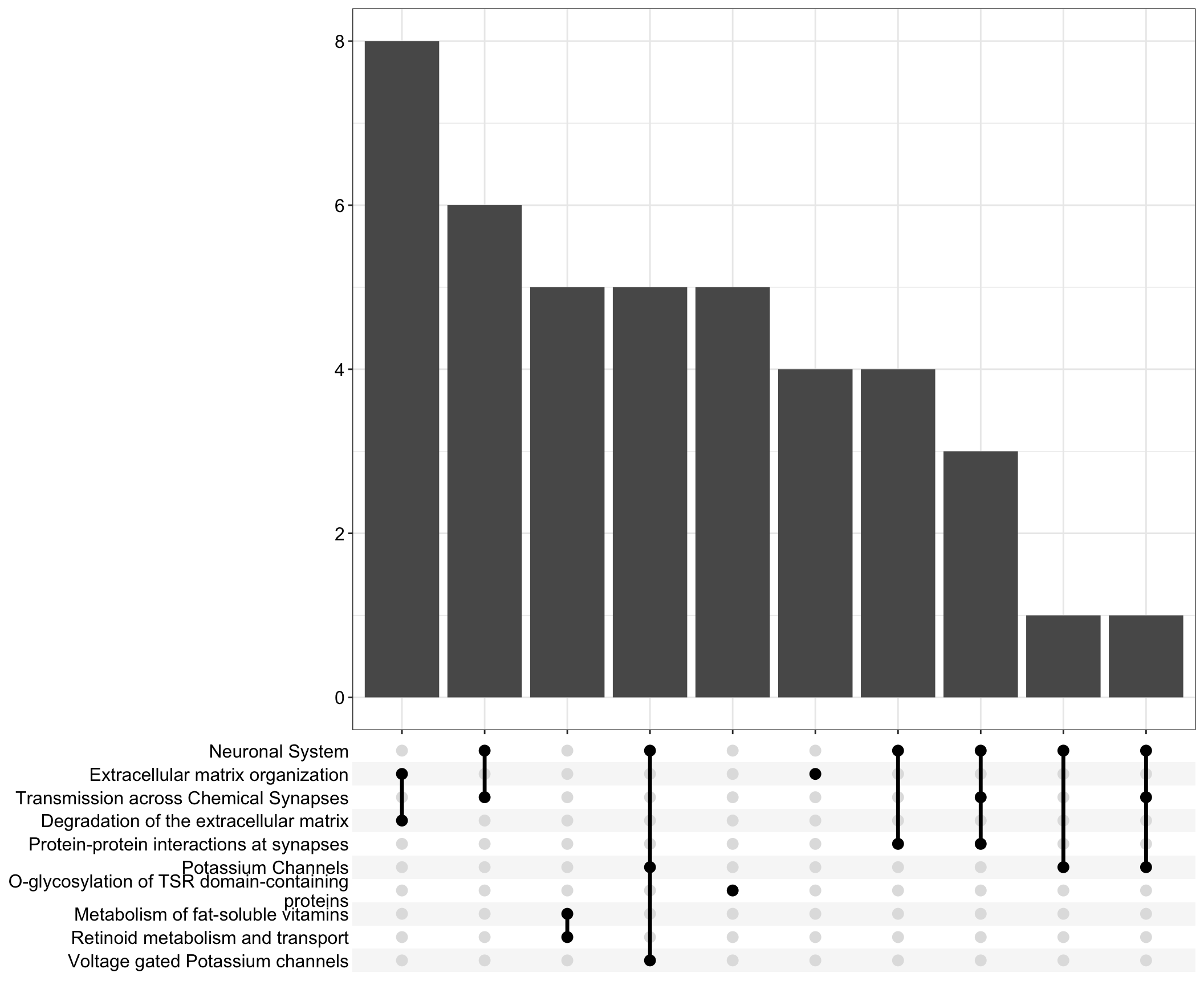
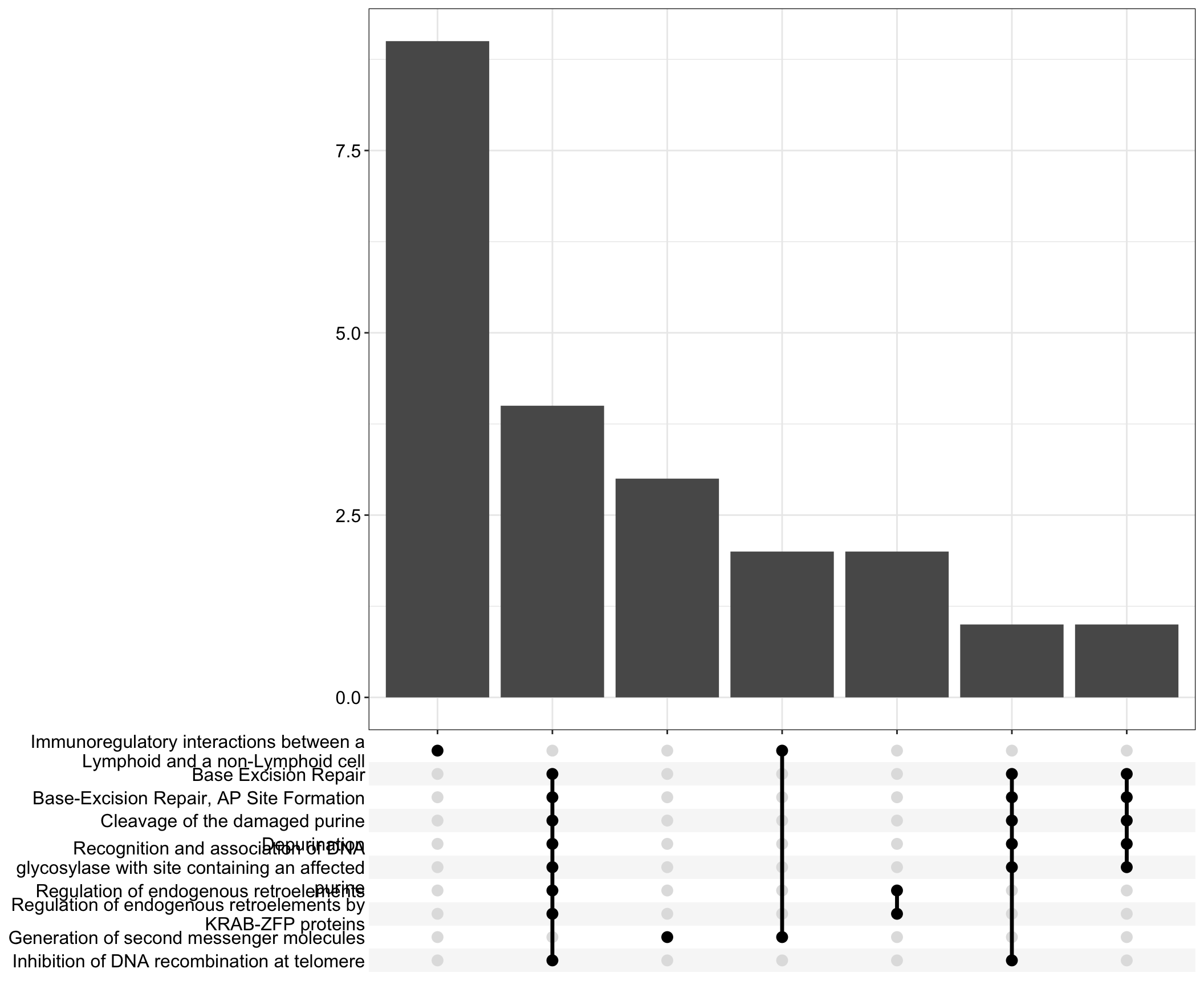
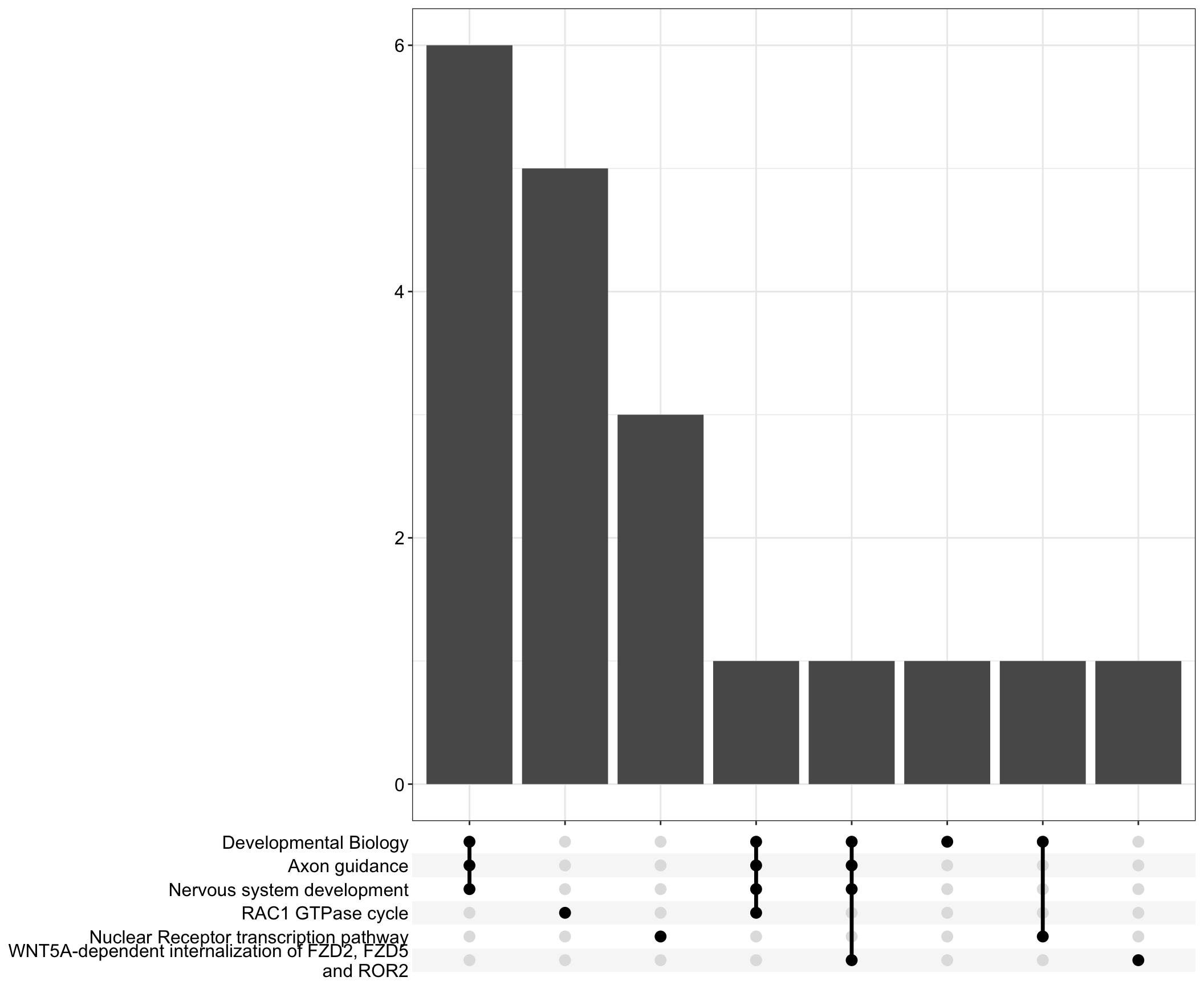
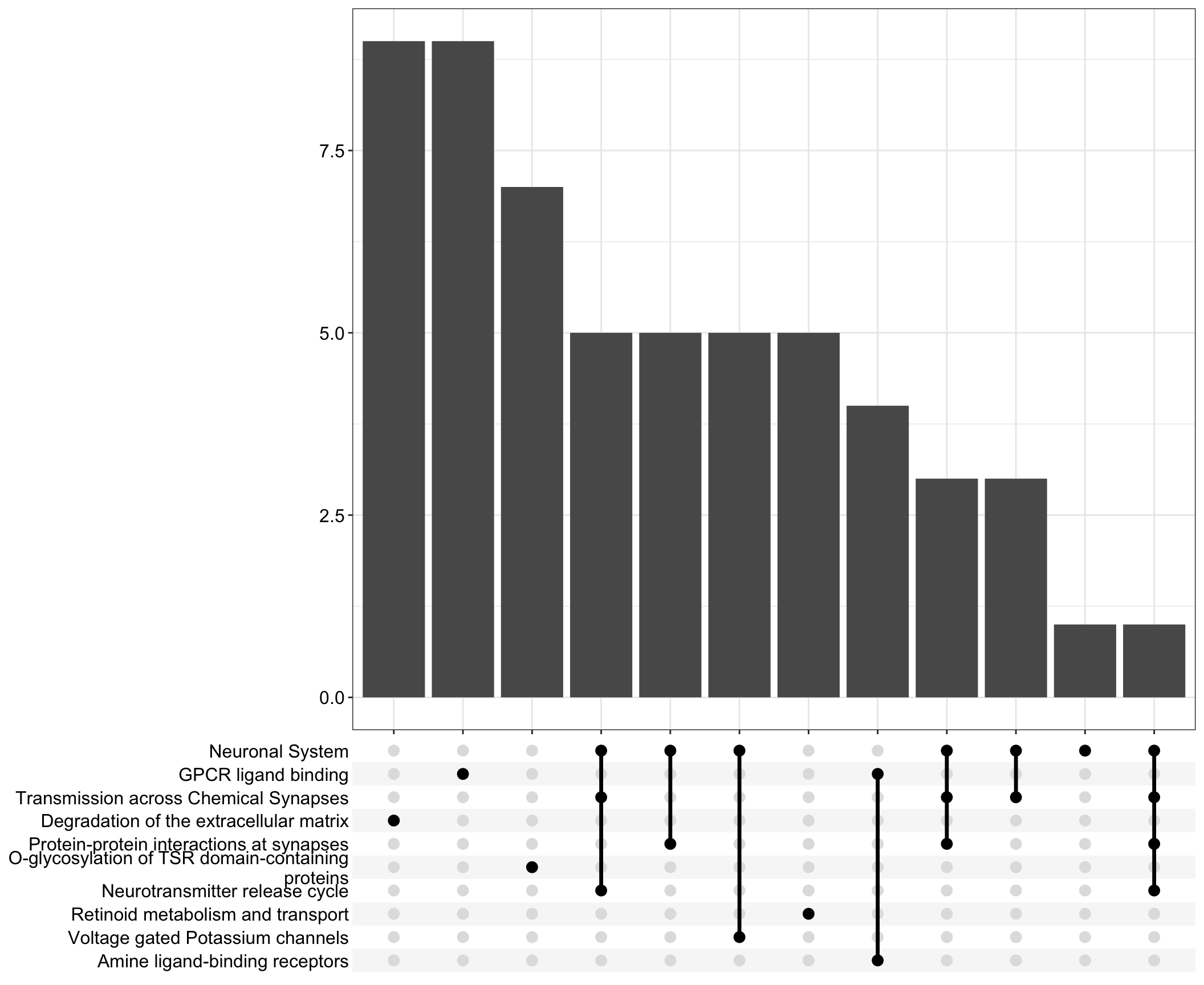
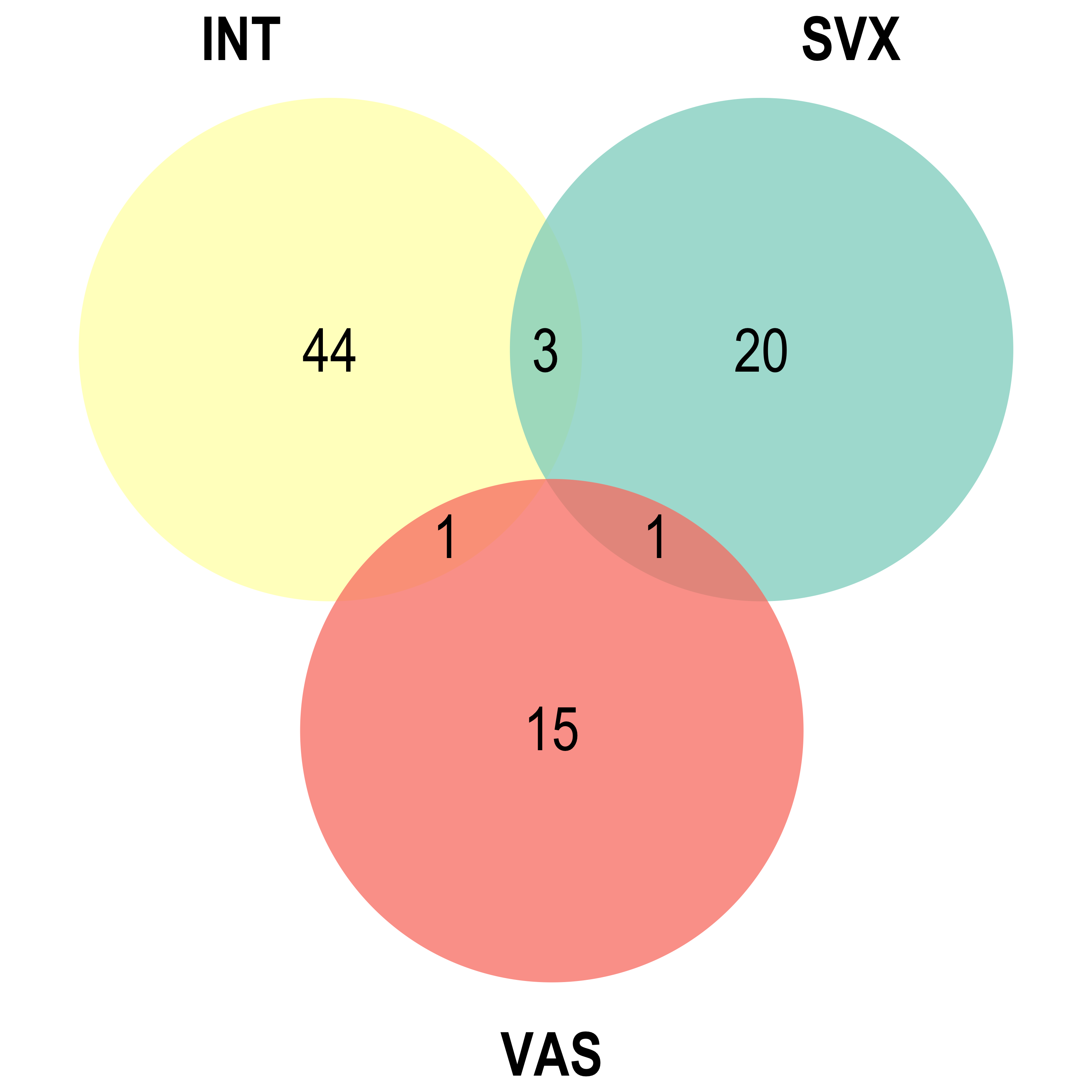
Upset: illustrate the overlap of gene between different pathways
I recommend reading through the full list of significant Reactome pathways and selecting the most biologically relevant for better visualisation
The following visualisations are Reactome enrichment analysis performed with set of DE genes. IMPORTANTLY, significant Reactome pathways are significantly if P value < 0.05
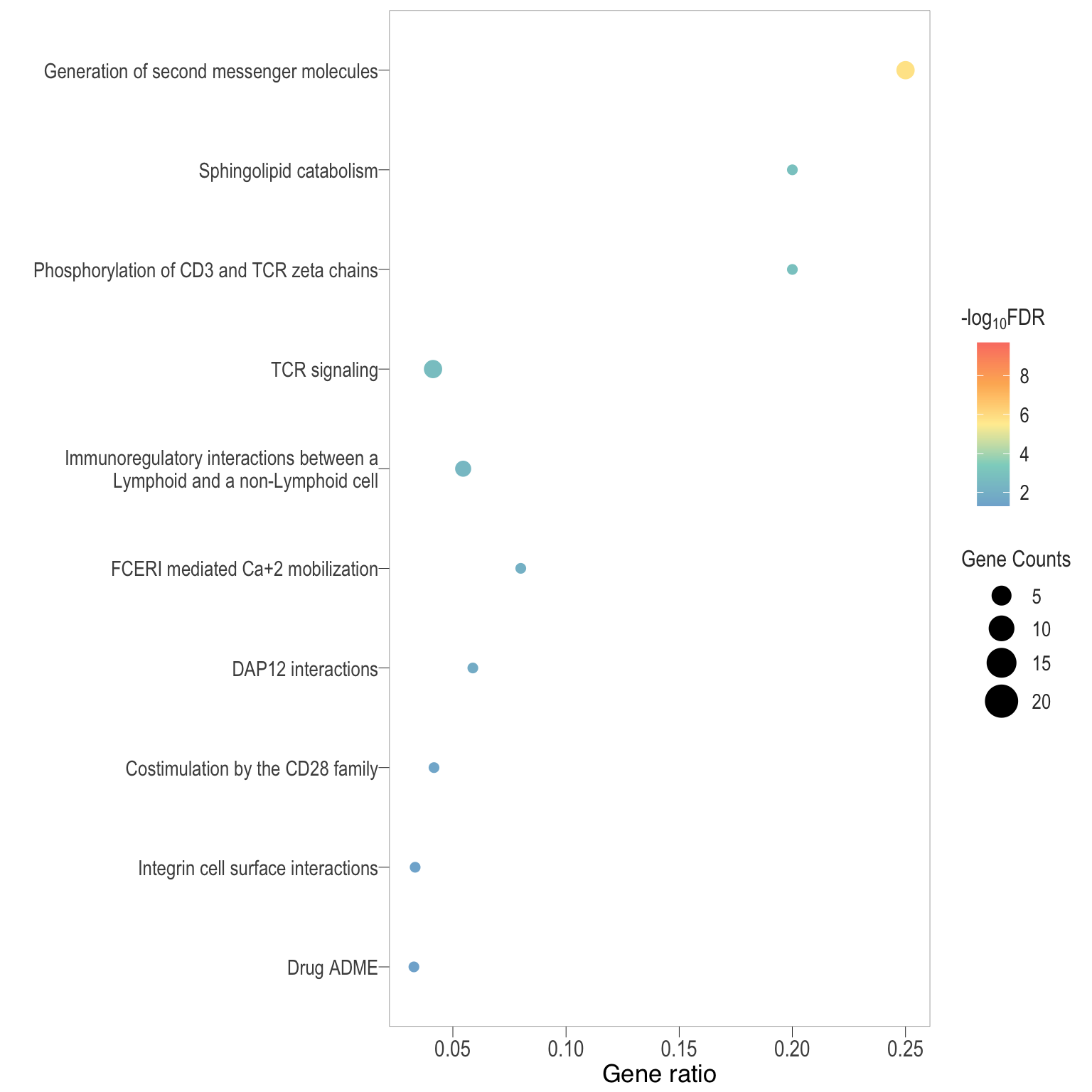
Dot plot: illustrates the enriched Reactome pathways
- \(Gene ratio =\) the number of significant DE gene in the term / the total of number of genes in the term as indicated by the size
Table: list of all the significant Reactome pathways
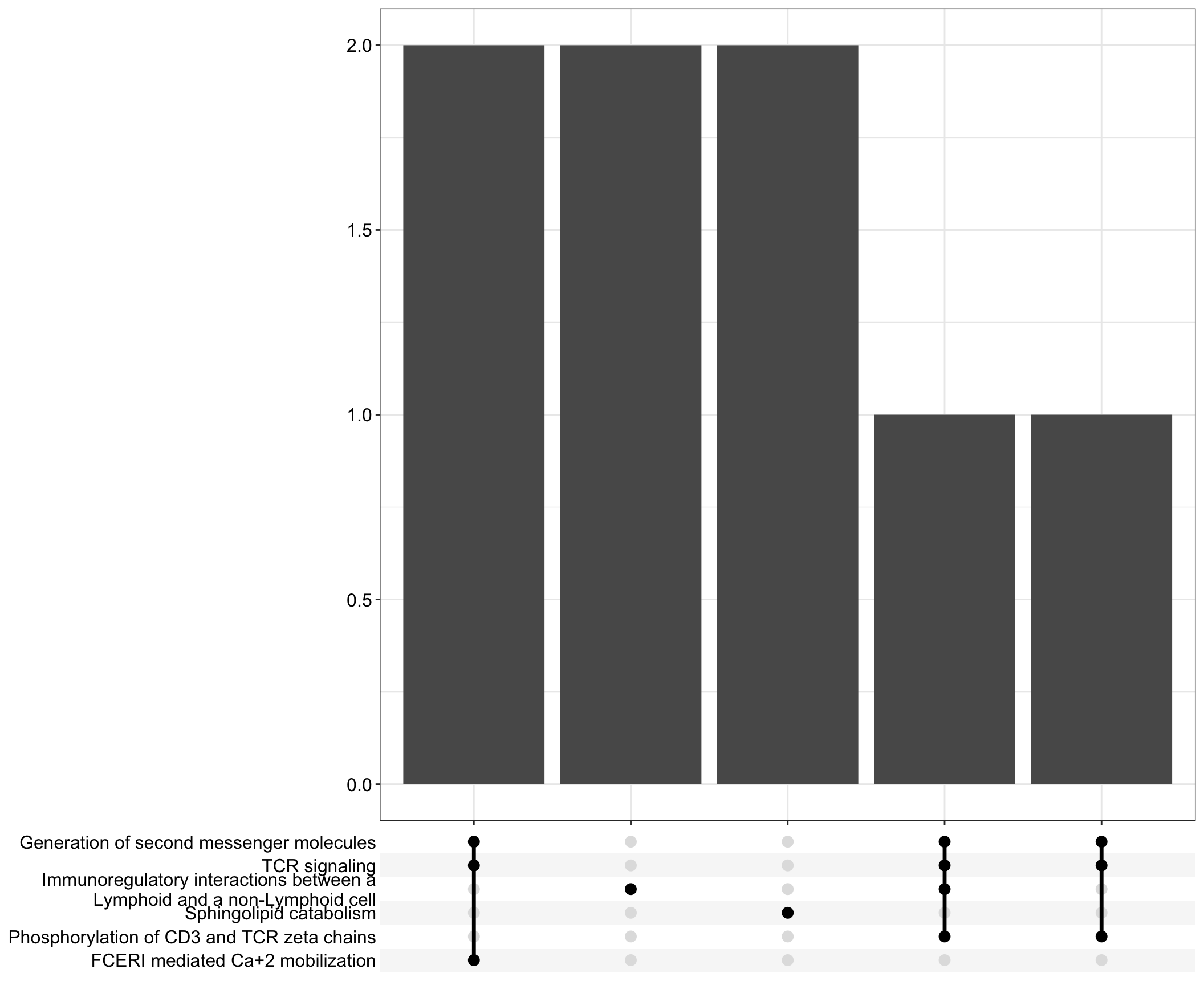
Upset: illustrate the overlap of gene between different pathways
I recommend reading through the full list of significant Reactome pathways and selecting the most biologically relevant for more in-depth visualisation
INT vs CONT
Table
INT vs SVX_VAS
Table
SVX vs SVX_VAS
Table
VAS vs SVX_VAS
Table
SVX_VAS vs CONT
Table
INT vs VAS
Table
Export Data
R version 4.4.1 (2024-06-14)
Platform: aarch64-apple-darwin20
Running under: macOS 26.1
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.0
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
time zone: Australia/Adelaide
tzcode source: internal
attached base packages:
[1] stats4 grid stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] knitr_1.50 pandoc_0.2.0 ReactomePA_1.50.0
[4] enrichplot_1.26.6 org.Mm.eg.db_3.20.0 AnnotationDbi_1.68.0
[7] IRanges_2.40.1 S4Vectors_0.44.0 Biobase_2.66.0
[10] BiocGenerics_0.52.0 clusterProfiler_4.14.6 Glimma_2.16.0
[13] edgeR_4.4.2 limma_3.62.2 ggrepel_0.9.6
[16] ggbiplot_0.6.2 extrafont_0.19 DT_0.34.0
[19] pheatmap_1.0.13 cowplot_1.2.0 pander_0.6.6
[22] kableExtra_1.4.0 VennDiagram_1.7.3 futile.logger_1.4.3
[25] lubridate_1.9.4 forcats_1.0.0 stringr_1.5.2
[28] purrr_1.1.0 tidyr_1.3.1 ggplot2_4.0.0
[31] tidyverse_2.0.0 reshape2_1.4.4 tibble_3.3.0
[34] readr_2.1.5 magrittr_2.0.4 dplyr_1.1.4
loaded via a namespace (and not attached):
[1] splines_4.4.1 later_1.4.4
[3] ggplotify_0.1.3 R.oo_1.27.1
[5] polyclip_1.10-7 graph_1.84.1
[7] lifecycle_1.0.4 rprojroot_2.1.1
[9] lattice_0.22-7 MASS_7.3-65
[11] crosstalk_1.2.2 sass_0.4.10
[13] rmarkdown_2.29 jquerylib_0.1.4
[15] yaml_2.3.10 httpuv_1.6.16
[17] ggtangle_0.0.7 DBI_1.2.3
[19] RColorBrewer_1.1-3 abind_1.4-8
[21] zlibbioc_1.52.0 GenomicRanges_1.58.0
[23] R.utils_2.13.0 ggraph_2.2.2
[25] yulab.utils_0.2.1 tweenr_2.0.3
[27] rappdirs_0.3.3 git2r_0.36.2
[29] GenomeInfoDbData_1.2.13 tidytree_0.4.6
[31] reactome.db_1.89.0 svglite_2.2.1
[33] codetools_0.2-20 DelayedArray_0.32.0
[35] DOSE_4.0.1 xml2_1.4.0
[37] ggforce_0.5.0 tidyselect_1.2.1
[39] aplot_0.2.9 UCSC.utils_1.2.0
[41] farver_2.1.2 viridis_0.6.5
[43] matrixStats_1.5.0 jsonlite_2.0.0
[45] tidygraph_1.3.1 systemfonts_1.2.3
[47] tools_4.4.1 ragg_1.5.0
[49] treeio_1.30.0 Rcpp_1.1.0
[51] glue_1.8.0 gridExtra_2.3
[53] Rttf2pt1_1.3.12 SparseArray_1.6.2
[55] here_1.0.2 xfun_0.53
[57] DESeq2_1.46.0 qvalue_2.38.0
[59] MatrixGenerics_1.18.1 GenomeInfoDb_1.42.3
[61] withr_3.0.2 formatR_1.14
[63] fastmap_1.2.0 digest_0.6.37
[65] timechange_0.3.0 R6_2.6.1
[67] gridGraphics_0.5-1 textshaping_1.0.3
[69] colorspace_2.1-1 GO.db_3.20.0
[71] RSQLite_2.4.3 R.methodsS3_1.8.2
[73] generics_0.1.4 data.table_1.17.8
[75] graphlayouts_1.2.2 httr_1.4.7
[77] htmlwidgets_1.6.4 S4Arrays_1.6.0
[79] graphite_1.52.0 whisker_0.4.1
[81] pkgconfig_2.0.3 gtable_0.3.6
[83] blob_1.2.4 workflowr_1.7.2
[85] S7_0.2.0 XVector_0.46.0
[87] htmltools_0.5.8.1 fgsea_1.32.4
[89] ggupset_0.4.1 scales_1.4.0
[91] png_0.1-8 ggfun_0.2.0
[93] lambda.r_1.2.4 rstudioapi_0.17.1
[95] tzdb_0.5.0 nlme_3.1-168
[97] cachem_1.1.0 parallel_4.4.1
[99] pillar_1.11.1 vctrs_0.6.5
[101] promises_1.3.3 extrafontdb_1.0
[103] evaluate_1.0.5 cli_3.6.5
[105] locfit_1.5-9.12 compiler_4.4.1
[107] futile.options_1.0.1 rlang_1.1.6
[109] crayon_1.5.3 labeling_0.4.3
[111] plyr_1.8.9 fs_1.6.6
[113] writexl_1.5.4 stringi_1.8.7
[115] viridisLite_0.4.2 BiocParallel_1.40.2
[117] Biostrings_2.74.1 lazyeval_0.2.2
[119] GOSemSim_2.32.0 Matrix_1.7-4
[121] hms_1.1.3 patchwork_1.3.2
[123] bit64_4.6.0-1 KEGGREST_1.46.0
[125] statmod_1.5.0 SummarizedExperiment_1.36.0
[127] igraph_2.1.4 memoise_2.0.1
[129] bslib_0.9.0 ggtree_3.14.0
[131] fastmatch_1.1-6 bit_4.6.0
[133] ape_5.8-1 gson_0.1.0